Approximately one to two out of every 100 newborns enter the world with a congenital heart defect, commonly abbreviated as CHD, but the precise origins of these conditions frequently remain shrouded in mystery. Genetic specialists from University Medicine Oldenburg in Germany have introduced an innovative approach to ascertain whether variations in the NOTCH1 gene are directly responsible for such defects. This breakthrough holds significant promise, as it empowers medical professionals to deliver more accurate diagnoses and craft precise, effective treatment strategies tailored to individual needs. Moreover, it offers families much-needed clarity on whether their child’s heart condition is passed down genetically, potentially influencing future health decisions and family planning.
The research team, under the leadership of Professor Dr. Marc-Phillip Hitz, who serves as the Director of the University Institute for Medical Genetics at Klinikum Oldenburg, along with Dr. Gregor Dombrowsky, the lead author of the study, detailed their discoveries in the prestigious journal Genome Medicine. Their work addresses a longstanding challenge in pediatric cardiology and genetics.
Identifying the specific trigger behind a congenital heart defect can feel akin to searching for a single needle amidst an enormous haystack. The human genome comprises roughly 20,000 genes, and within each gene, numerous variants can exist, with certain ones exerting detrimental influences on development. Frequently, these heart anomalies arise from a confluence of several genetic changes happening concurrently, creating an exponentially large array of possible interactions and combinations that complicate diagnosis.
In their comprehensive investigation, scientists from Oldenburg performed detailed molecular genetic examinations on blood samples collected from nearly 4,000 children diagnosed with heart conditions. Amidst this extensive dataset, one gene repeatedly emerged as a prominent factor: NOTCH1. This gene is responsible for producing a critical signaling protein that is essential during the embryonic stage for properly shaping and forming the heart.
When alterations occur in the genetic blueprint provided by NOTCH1, the signaling protein it produces becomes dysfunctional. This malfunction then interferes with the intricate and delicate molecular processes that govern heart development in the embryo. Even a minor deviation in this blueprint can initiate a chain reaction of developmental errors, culminating in serious structural abnormalities known as congenital heart defects. While mutations in NOTCH1 are implicated in only about 1% of all CHD cases, it stands out as the leading single-gene culprit among monogenic causes of these defects.
Dr. Dombrowsky elaborates, “Certain variants within this gene have long been recognized as contributors to congenital heart defects, often identified after excluding alternative explanations. However, in clinical settings, we regularly encounter a wide array of NOTCH1 variants, including many novel ones, and it remains uncertain whether they are benign or truly pathogenic.” To bridge this knowledge gap, the research team sought a reliable method to evaluate the impact of these variants definitively.
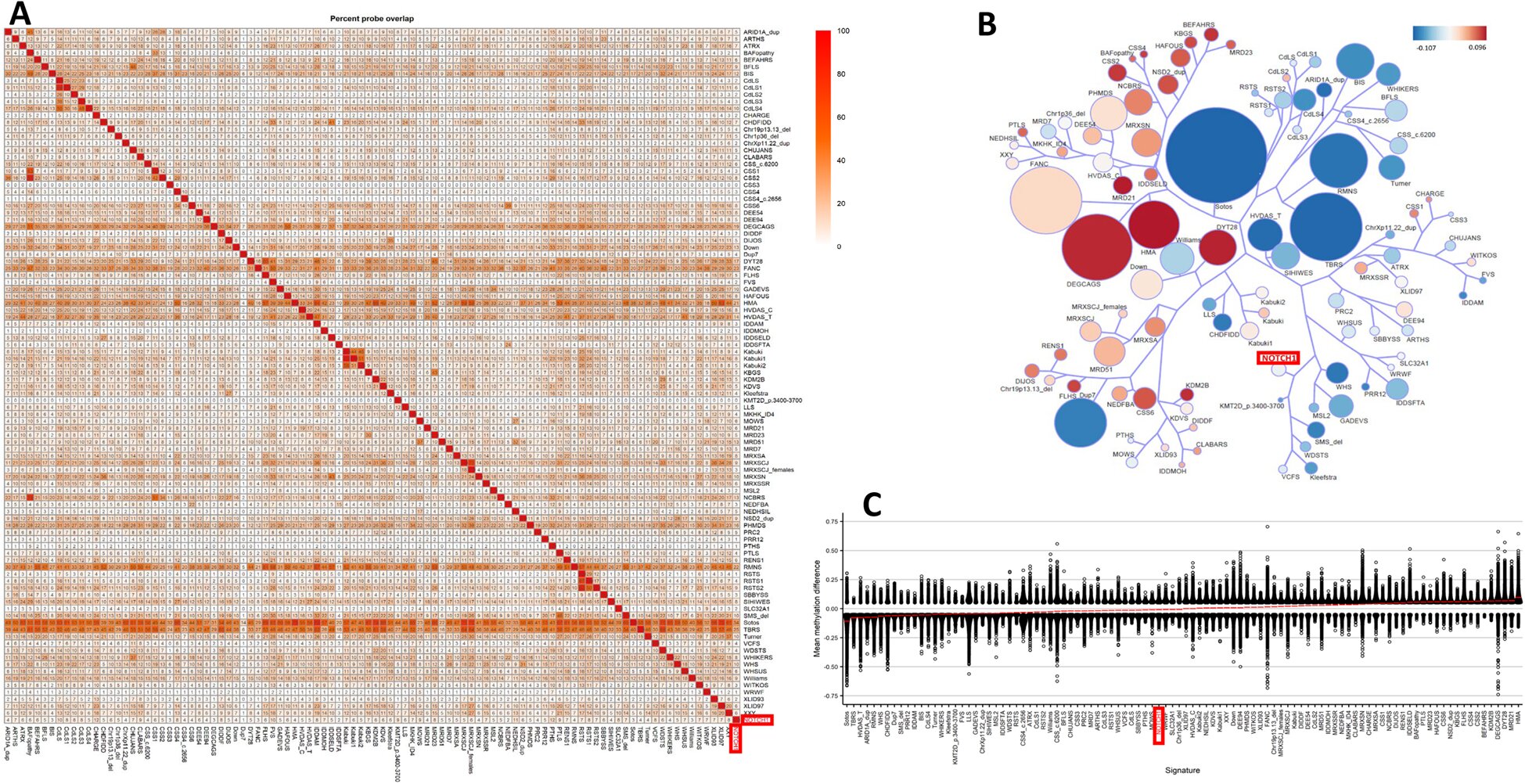
Approaching their task with the meticulousness of detectives unraveling a complex case, the researchers traced the molecular signatures left by established, severe NOTCH1 variants in the DNA of patients affected by heart defects. They understood that the signaling disruption triggered by these genetic flaws ultimately impairs the proper transcription of other genes that are structurally normal. A key mechanism involved here is DNA methylation, where methyl groups bind to specific sites along the DNA strand.
The investigators hypothesized that in individuals carrying pathogenic NOTCH1 variants, these methylation patterns would exhibit distinct alterations compared to healthy controls. Through rigorous analysis, the Oldenburg team uncovered a remarkable consistency: across patients harboring various NOTCH1 variants, identical genomic regions displayed modified methylation profiles.
Dr. Dombrowsky describes this finding vividly: “The consistent pattern of these methylation changes across affected DNA segments acts like a unique fingerprint imprinted by a pathogenic NOTCH1 variant on the genome. Armed with this signature, clinicians can now confidently determine if a specific variant is driving a patient’s heart defect, revolutionizing diagnostic accuracy.”
This discovery not only enhances diagnostic precision but also opens doors to personalized medicine in the realm of congenital heart diseases. For families, it means moving from uncertainty to informed understanding—knowing if the condition is hereditary can guide genetic counseling, prenatal screening in future pregnancies, and even long-term monitoring for related health risks in siblings or other relatives.
The complexity of congenital heart defects stems from their multifactorial nature. Environmental influences, such as maternal diabetes, infections during pregnancy, or exposure to certain medications, can interact with genetic predispositions. However, when a clear monogenic cause like NOTCH1 is identified, it simplifies the picture and allows for targeted interventions. For instance, early identification might prompt closer echocardiographic surveillance or even experimental therapies aimed at modulating the NOTCH signaling pathway.
The NOTCH1 protein’s role in development is profound. It belongs to a family of receptors that influence cell fate decisions during embryogenesis, particularly in tissues forming the cardiovascular system, valves, and septa of the heart. Disruptions here can lead to conditions like bicuspid aortic valve, hypoplastic left heart syndrome, or tetralogy of Fallot, each requiring specialized surgical corrections.
The methylation episignature—the term for this distinctive pattern—represents a paradigm shift. Unlike direct sequencing, which identifies variants but not their functional impact, this epigenetic readout provides functional evidence of pathogenicity. It’s particularly valuable for variants of uncertain significance (VUS), which plague genetic testing panels today.
In practical terms, implementing this test could integrate into routine clinical workflows. A blood sample from a newborn with CHD, sequenced for NOTCH1 variants, followed by methylation profiling, could yield results within weeks, informing surgical planning and family discussions. As sequencing costs drop and epigenetic assays become standardized, widespread adoption seems feasible.
Looking ahead, this methodology might extend beyond NOTCH1 to other CHD genes, building a comprehensive episignature library for non-syndromic heart defects. Collaborations between Oldenburg’s team and international consortia could validate and refine these signatures across diverse populations, addressing genetic heterogeneity.
For patients and families, the emotional weight of a CHD diagnosis is immense. Uncertainty about heritability amplifies anxiety—will it affect future children? This test alleviates that by providing concrete data, fostering empowerment through knowledge. It underscores how advances in epigenetics are transforming rare disease diagnostics, making precision medicine accessible even for conditions affecting 1-2% of births globally.
The study’s rigorous design, involving nearly 4,000 samples, lends robust statistical power. Controls without CHD helped delineate disease-specific patterns, while comparisons to other syndromes ensured specificity. Published in Genome Medicine, it underwent peer review, affirming its scientific merit.
Publication Details
The full study, titled “DNA methylation analysis of NOTCH1 variants reveals the first episignature for non-syndromic congenital heart defects,” was authored by Gregor Dombrowsky and colleagues. It appeared in Genome Medicine in 2026, with DOI: 10.1186/s13073-025-01587-6.